Page 42 - 《南京医科大学学报》自然科学版2026年第2期
P. 42
第46卷第2期
·198 · 南 京 医 科 大 学 学 报 2026年2月
A B hAMSC⁃exo⁃F hAMSC⁃exo⁃M
Group
hAMSC⁃exo⁃F vs.hAMSC⁃exo⁃M hsa⁃miR⁃335⁃3p 1.5
1
hsa⁃miR⁃301a⁃3p 1.Glycerophospholipid metabolism
hsa⁃miR⁃367⁃3p 0.5
hsa⁃miR⁃1268b
hsa⁃miR⁃664a⁃5p Glycosaminoglycan biosynthesis
0
hsa⁃miR⁃149⁃5p
hsa⁃miR⁃34a⁃5p ⁃0.5
hsa⁃miR⁃1291 Arachidonic acid metabolism
hsa⁃miR⁃218⁃5p
3 hsa⁃miR⁃218⁃5p ⁃1
hsa⁃miR⁃29b⁃3p hsa⁃miR⁃29a⁃3p ⁃1.5 C hAMSC⁃exo⁃F
hsa⁃miR⁃708⁃5p
hsa⁃miR⁃1291
hsa⁃miR⁃136⁃3p
hsa⁃miR⁃486⁃5p
P 2 Up hsa⁃miR⁃4454
-lg hsa⁃miR⁃328⁃3p hsa⁃miR⁃34a⁃5p Down hsa⁃miR⁃486⁃5p
hsa⁃miR⁃664a⁃5p Normal
hsa⁃miR⁃584⁃5p
71 337 50
hsa⁃miR⁃671⁃5p
1
hsa⁃miR⁃92a⁃3p
hsa⁃miR⁃328⁃3p
hsa⁃miR⁃1268a
hsa⁃miR⁃1268b
0 2.ECM⁃receptor interaction
hsa⁃miR⁃193b⁃5p hAMSC⁃exo⁃M
-5.0 -2.5 0 2.5 hsa⁃miR⁃149⁃5p Focal adhesion
log2 FC hsa⁃miR⁃362⁃5p Glutamatergic synapse
hsa⁃miR⁃708⁃5p
Pathways in cancer
hsa⁃miR⁃1287⁃5p
Inflammatory mediator regula⁃
hsa⁃miR⁃377⁃3p
hsa⁃miR⁃2682⁃3p tion of TRP channels
hsa⁃miR⁃484
hsa⁃miR⁃3182
hsa⁃miR⁃3615
D 0 2 4 6 8 E 0 2 4 6 8
Pathways in cancer cAMP signaling pathway
Adrenergic signaling in cardiac myocytes Focal adhesion
Focal adhesion Proteoglycans in cancer
Proteoglycans in cancer Endocytosis
MAPK signaling pathway Insulin secretion
hAMSC⁃exo⁃F unique Estrogen signaling pathway hAMSC⁃exo⁃M unique Rap1 signaling pathway
Cholinergic synapse
Rap1 signaling pathway
Circadian entrainment
Insulin secretion
Adrenergic signaling in cardiomyocytes
cGMP⁃PKG signaling pathway
Small cell lung cancer
ECM⁃receptor interaction
Wnt signaling pathway
Glycerophospholipid metabolism
Signaling pathways regulating pluripotency of stem cells
Non⁃small cell lung cancer
Neurotrophin signaling pathway
Pl3K⁃Akt signaling pathway
ECM⁃receptor interaction
Choline metabolism in cancer
Hedgehog signaling pathway
HIF⁃1 signaling pathway
Oxytocin signaling pathway Pathways in cancer
cAMP signaling pathway Melanogenesis
Axon guidance Basal cell carcinoma
VEGF signating pathway MAPK signaling pathway
Toxoplasmosis Wnt signaling pathway
0 2 4 6 8 0 2 4 6 8
-lg P -lg P
F
0 2 4 6 8
Dilated cardiomyopathy
Estrogen signaling pathway
Pathways in cancer
Focal adhesion
Endocytosis
Platelet activation
ECM⁃receptor interaction
Rap1 signaling pathway
Common Melanogensis
Thyroid hormone signaling pathway
Axon guidance
Arrhythmogenic right ventricular cardiomyopathy(ARVC)
Calclum signaling pathway
Proteoglycans in cancer
Non⁃small cell lung cancer
Basal cell carcinoma
Signaling pathways regulating pluripotency of stem cells
Adrenergic signaling in eardiomyocytes
TypeⅡdiabetes mellitus
Hedgehog signaling pathway
0 2 4 6 8
-lg P
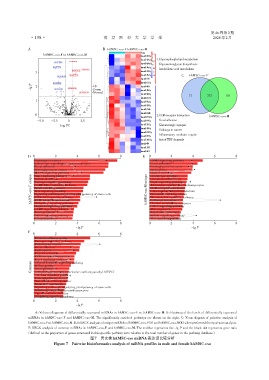
A:Volcano diagrams of differentially expressed miRNAs in hAMSC⁃exo⁃F vs. hAMSC⁃exo⁃M. B:Heatmap of the levels of differentially expressed
miRNAs in hAMSC⁃exo⁃F and hAMSC⁃exo⁃M. The significantly enriched pathways are shown on the right. C:Venn diagram of pairwise analysis of
hAMSC⁃exo⁃Fvs.hAMSC⁃exo⁃M.D,E:KEGGanalysisofuniquemiRNAsofhAMSC⁃exo⁃F(D)andhAMSC⁃exo⁃M(E)whenperformedtheirpairwiseanalysis.
F:KEGG analysis of common miRNAs in hAMSC⁃exo⁃F and hAMSC⁃exo⁃M. The red bar represents the ⁃lg P and the black dot represents gene ratio
(defined as the proportion of genes annotated in this specific pathway term relative to the total number of genes in the pathway database).
图7 男女株hAMSC⁃exo miRNA表达谱比较分析
Figure 7 Pairwise bioinformatics analysis of miRNA profiles in male and female hAMSC⁃exo