Page 41 - 《南京医科大学学报》自然科学版2026年第2期
P. 41
第46卷第2期 黄蓓蓓,宁 松,吴慧敏,等. 不同性别来源人胚胎干细胞与羊膜间充质干细胞外泌体蛋白质组学与
2026年2月 miRNA谱分析[J]. 南京医科大学学报(自然科学版),2026,46(2):188-201 ·197 ·
A B hESC⁃exo⁃F hESC⁃exo⁃M
Group
hsa⁃miR⁃574⁃5p
hsa⁃miR⁃125b⁃5p 1
hESC⁃exo⁃F vs. hESC⁃exo⁃M hsa⁃miR⁃1268a 1. Cholinergic synapse
hsa⁃miR⁃1268b
hsa⁃miR⁃1275 0
hsa⁃miR⁃4516
hsa⁃let⁃7b⁃5p ⁃1 Focal adhesion
hsa⁃miR⁃503⁃5p
hsa⁃miR⁃367⁃3p
hsa⁃let⁃7a⁃5p
hsa⁃let⁃7c⁃5p Oxytocin signaling pathway
hsa⁃miR⁃152⁃3p
hsa⁃miR⁃27a⁃3p
7.5 hsa⁃miR⁃27b⁃3p MAPK signaling pathway C hESC⁃exo⁃F
hsa⁃miR⁃23a⁃3p
hsa⁃miR⁃23b⁃3p
hsa⁃let⁃7e⁃5p
hsa⁃let⁃7f⁃5p
hsa⁃miR⁃145⁃5p Axon guidance
hsa⁃miR⁃483⁃5p
hsa⁃miR⁃4791
hsa⁃miR⁃484 hsa⁃miR⁃28⁃5p
hsa⁃miR⁃143⁃3p
hsa⁃miR⁃210⁃3p hsa⁃miR⁃3679⁃5p
hsa⁃miR⁃374b⁃3p
P 5.0 hsa⁃miR⁃1180⁃3p Up hsa⁃miR⁃708⁃5p
hsa⁃miR⁃26a⁃5p
hsa⁃miR⁃302c⁃3p
-lg hsa⁃miR⁃425⁃5p Down hsa⁃miR⁃454⁃3p 179 377 35
hsa⁃miR⁃372⁃3p
hsa⁃miR⁃484
hsa⁃miR⁃4516 Normal hsa⁃miR⁃181a⁃5p
hsa⁃miR⁃1287⁃5p
hsa⁃miR⁃378g
hsa⁃miR⁃23a⁃3p
hsa⁃miR⁃574⁃5p hsa⁃miR⁃378d
hsa⁃miR⁃378f
2.5 hsa⁃miR⁃302d⁃5p
hsa⁃miR⁃378e
hsa⁃miR⁃3679⁃5p hsa⁃miR⁃301b⁃3p
hsa⁃miR⁃27b⁃3p hsa⁃miR⁃877⁃3p
hsa⁃miR⁃1277⁃5p
hsa⁃miR⁃1180⁃3p
hsa⁃miR⁃367⁃3p
hsa⁃miR⁃1306⁃5p
hsa⁃miR⁃371a⁃5p
hsa⁃miR⁃378c hESC⁃exo⁃M
hsa⁃miR⁃378a⁃3p
hsa⁃miR⁃378i 2. Type II diabetes mellitus
0 hsa⁃miR⁃769⁃5p
hsa⁃miR⁃17⁃3p
hsa⁃miR⁃296⁃5p Focal adhesion
-3 0 3 6 hsa⁃miR⁃106b⁃3p
hsa⁃miR⁃320a⁃3p
hsa⁃miR⁃320b
hsa⁃miR⁃328⁃3p Glutamatergic synapse
log2 FC hsa⁃miR⁃92b⁃3p
hsa⁃miR⁃3615
hsa⁃miR⁃92a⁃3p Inflammatory mediator regula⁃
hsa⁃miR⁃423⁃3p
hsa⁃miR⁃210⁃3p
hsa⁃miR⁃486⁃5p
hsa⁃miR⁃301a⁃3p tion of TRP channels
hsa⁃miR⁃342⁃3p
hsa⁃miR⁃219a⁃5p
hsa⁃miR⁃378h
hsa⁃miR⁃16⁃2⁃3p Platelet activation
hsa⁃miR⁃548f⁃3p
hsa⁃miR⁃338⁃3p
hsa⁃miR⁃548g⁃3p
hsa⁃miR⁃363⁃3p
hsa⁃miR⁃200a⁃3p
hsa⁃miR⁃425⁃5p
hsa⁃miR⁃1296⁃5p
hsa⁃miR⁃124⁃3p
hsa⁃miR⁃32⁃5p
hsa⁃miR⁃135a⁃5p
hsa⁃miR⁃135b⁃5p
hsa⁃miR⁃200c⁃3p
hsa⁃miR⁃218⁃5p
hsa⁃miR⁃103a⁃3p
hsa⁃miR⁃107
0 2.5 5.0 7.5 10.0 0 2 4 6 8 10
D E
Focal adhesion cAMP signaling pathway
ECM⁃receptor interaction Oxytocin signaling pathwau
cAMP signaling pathway MAPK signaling pathway
Axon guidace Pathways in cancer
MAPK signaling pathway Rap1 signaling pathway
Dilated cardiomyopathy Ras signaling pathway
hESC⁃exo⁃F unique Ras signaling pathway hESC⁃exo⁃M unique Adrenergic signaling in cardiomyocytes
PI3K⁃Akt signaling pathway
Wnt signaling pathway
ErbB signaling pathway
Rap1 signaling pathway
Sphingolipid signaling pathway
Adrenergic signaling in cardiomyocytes
Cholinergic synapse
FoxO signaling pathway
Chronic myeloid leukemia
MicroRNAs in cancer
Tight junction
Glutamatergic synapce
Insulin signaling pathway
Hypertrophic car diomyopathy(HCM)
Neurotrophin signaling pathway
Oxytocin signaling pathway HTLV⁃ infection
Inflammatory mediator regulation of TRP channels
Arrhythmogeric right ventricular cardiomyopathy(ARVC) Adherens junction
Circadian entrairment Proteoglycans in cancer
Notch signaling pathway Calcium signaling pathway
Retrograde endocannabinoid signaling Thyroid hormone signaling pathway
0 2.5 5.0 7.5 10.0 0 2 4 6 8 10
-lg P -lg P
F 0 2 4 6
Focal adhesion
Pathways in cancer
Endocytosis
ECM⁃receptor interaction
Staphylococcuc aureus infection
Signaling pathways regulating pluripotency of stem cells
AMPK signaling pathway
Rap1 signaling pathway
Common Cholinergic synapse
Ptoteoglycans in cancer
Non⁃small cell lung cancer
Estrogen signaling pathway
Thyroid hormone signaling pathway
Calcium signaling pathway
Prostate cancer
Toxoplasmosis
MicroRNAs in cancer
Acute myeloid leukemia
Bladder cancer
Hippo signaling pathway
0 2 4 6
-lg P
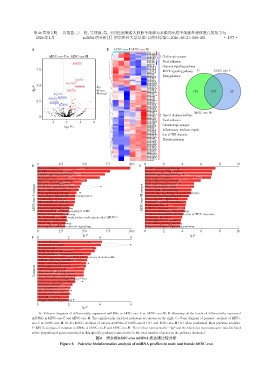
A:Volcano diagrams of differentially expressed miRNAs in hESC⁃exo⁃F vs. hESC⁃exo⁃M. B:Heatmap of the levels of differentially expressed
miRNAs in hESC⁃exo⁃F and hESC⁃exo⁃M. The significantly enriched pathways are shown on the right. C:Venn diagram of pairwise analysis of hESC⁃
exo⁃F vs. hESC⁃exo⁃M. D,E:KEGG analysis of unique miRNAs of hESC⁃exo⁃F(D)and hESC⁃exo⁃M(E)when performed their pairwise analysis.
F:KEGG analysis of common miRNAs in hESC⁃exo⁃F and hESC⁃exo⁃M. The red bar represents the -lgP and the black dot represents gene ratio(defined
as the proportion of genes annotated in this specific pathway term relative to the total number of genes in the pathway database).
图6 男女株hESC⁃exo miRNA表达谱比较分析
Figure 6 Pairwise bioinformatics analysis of miRNA profiles in male and female hESC⁃exo