Page 22 - 南京医科大学自然版
P. 22
第45卷第11期
·1552 · 南 京 医 科 大 学 学 报 2025年11月
A
shNC shKIF11⁃1
HCT116
10 Q1 Q2 10 Q1 Q2 50 ***
5
5
3.10% 7.41% 4.03% 15.4%
(%) 40
4
4
10 10
3 3 30
10 10 Apoptotic rate 20
10 2 10 2
Q4 Q3 Q4 Q3 10
0 0
-10 2 78.9% 10.6% -10 2 51.5% 29.1% 0
0 10 3 10 4 10 5 0 10 3 10 4 10 5 shNC shKIF11⁃1
5
5
10 Q1 Q2 10 Q1 Q2 50 DLD1
***
3.09% 9.06% 4.86% 16.2%
(%) 40
4
4
10 10
10 3 10 3 Apoptotic rate 30
20
10 2 10 2
Q4 Q3 Q4 Q3 10
0 0
-10 2 77.9% 9.94% -10 2 51.0% 27.9% 0
0 10 3 10 4 10 5 0 10 3 10 4 10 5 shNC shKIF11⁃1
B oe⁃NC oe⁃KIF11
5
5
10 Q1 Q2 10 Q1 Q2 15 SW480
0.41% 6.90% 0.27% 3.72% **
4 4 (%)
10 10 10
10 3 10 3 Apoptotic rate
10 2 10 2 5
Q4 Q3 Q4 Q3
0 0
88.2% 4.51% 92.2% 3.81%
0
3 4 5 3 4 5
0 10 10 10 0 10 10 10 oe⁃NC oe⁃KIF11
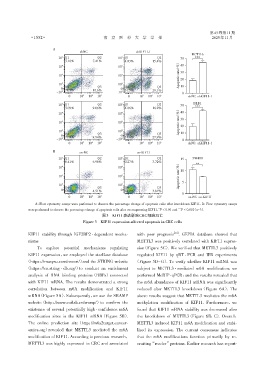
A:Flow cytometry assays were performed to observe the percentage change of apoptosis cells after knockdown KIF11. B:Flow cytometry assays
**
were performed to observe the percentage change of apoptosis cells after overexpressing KIF11. P < 0.01 and *** P < 0.001(n=3).
图3 KIF11表达影响CRC细胞凋亡
Figure 3 KIF11 expression affected apoptosis in CRC cells
[10]
KIF11 stability through IGF2BP2 ⁃ dependent mecha⁃ with poor prognosis . GEPIA database showed that
nisms METTL3 was positively correlated with KIF11 expres⁃
To explore potential mechanisms regulating sion(Figure 5C). We verified that METTL3 positively
KIF11 expression,we employed the starBase database regulated KIF11 by qRT ⁃ PCR and WB experiments
(https://rnasysu.com/encori/)and the STRING website (Figure 5D-G). To verify whether KIF11 mRNA was
(https://cn.string ⁃ db.org/)to conduct an enrichment subject to METTL3 ⁃ mediated m6A modification,we
analysis of RNA binding proteins(RBPs)connected performed MeRIP⁃qPCR,and the results revealed that
with KIF11 mRNA. The results demonstrated a strong the m6A abundance of KIF11 mRNA was significantly
correlation between m6A modification and KIF11 reduced after METTL3 knockdown(Figure 6A). The
mRNA(Figure 5A). Subsequently,we use the SRAMP above results suggest that METTL3 mediates the m6A
website(http://www.cuilab.cn/sramp/)to confirm the methylation modification of KIF11. Furthermore,we
existence of several potentially high ⁃ confidence m6A found that KIF11 mRNA stability was decreased after
modification sites in the KIF11 mRNA(Figure 5B). the knockdown of METTL3(Figure 6B,C). Overall,
The online prediction site(http://m6a2target.cancer⁃ METTL3 induced KIF11 m6A modification and stabi⁃
omics.org)revealed that METTL3 mediated the m6A lized its expression. The current consensus indicates
modification of KIF11. According to previous research, that the m6A modifications function primarily by re⁃
METTL3 was highly expressed in CRC and associated cruiting“reader”proteins. Earlier research has report⁃