Page 24 - 南京医科大学自然版
P. 24
第45卷第11期
·1554 · 南 京 医 科 大 学 学 报 2025年11月
(%) Biological process
A 120 GO enrichment
Molecular function
Percent of genes 80
Cellular component
40
0 ⁃methy lation in response to stress translational initiation Pre⁃miRNA binding Pre⁃mRNA binding siRNA binding Dendritiv filoppodium Dendritic spine neck factor complex Perichromatin fibrils Box C/D RNP complex Paraspeckles
Messenger ribonucleoprotein comlpex assembly Obsolete positive regulation of telomerase RNA reverse transcriptase activity (guanine⁃N7) tRNA mRNA alternative polyadenylation Global gene silencing by mRNA cleavage pre⁃mRNA cleavage required for polyadenylation Endothelial to hematopoietic transition Regulation of mRNA stability involved Positive regulation of cap⁃independent Regulation of Schwann cell differentiation C5⁃methylcytidine⁃containing RNA binding Regula
B Prediction score distribution along the query sequence C RNA polymerase P < 0.001
1.0 TPM) 5 4 R=0.42
0.9
Combined soore 0.8 High confidence (KIF11 3 2
0.7 Very high confidence
0.6
Low confidence
0.5 Moderate confidence log2 1
0.4 2 3 4 5
0 1 000 2 000 3 000 4 000 5 000 log2 (METTL3 TPM)
Position
D E
1.5 ** *** shNC HCT116 DLD1 1.5 * ** shNC
shMETTL3
shMETTL3
Relative KIF11 mRNA expression 1.0 KIF11 shNC shMETTL3 shNC shMETTL3 Relative KIF11 protein expression 1.0
130 kDa
0.5
0.5
0 ACTIN 42 kDa 0
HCT116 DLD1 HCT116 DLD1
F 5 4 *** * oe⁃NC G HCT116 DLD1 1.5 * * oe⁃NC
oe⁃METTL3
oe⁃METTL3
Relative KIF11 mRNA expression 3 2 KIF11 oe⁃NC oe⁃METTL3 oe⁃METTL3 Relative KIF11 protein expression 1.0
oe⁃NC
130 kDa
0.5
0 1 ACTIN 42 kDa 0
HCT116 DLD1 HCT116 DLD1
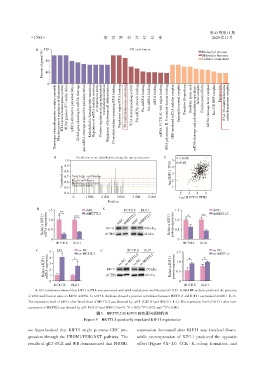
A:GO enrichment showed that KIF11 mRNA was associated with m6A methylation modification(n=118). B:SRAMP website predicted the presence
of m6A modification sites on KIF11 mRNA. C:GEPIA database showed a positive correlation between METTL3 and KIF11 expression(n=401). D,E:
The expression level of KIF11 after knockdown of METTL3 was detected by qRT⁃PCR(D)and WB(E). F,G:The expression level of KIF11 after over⁃
*
**
expression of METTL3 was dtected by qRT⁃PCR(F)and WB(G)(n=3). P < 0.05,P < 0.01,and *** P < 0.001.
图5 METTL3对KIF11具有正向调控作用
Figure 5 METTL3 positively regulated KIF11 exptression
we hypothesized that KIF11 might promote CRC pro⁃ expression decreased after KIF11 was knocked down,
gression through the PROM1/PI3K/AKT pathway. The while overexpression of KIF11 produced the opposite
results of qRT⁃PCR and WB demonstrated that PROM1 effect(Figure 9A- D). CCK ⁃ 8,colony formation,and