Page 67 - 《南京医科大学学报》2026年第1期
P. 67
第46卷第1期 马 金,沈 滟,沈国民,等. F9基因剪接位点共识区域中内含子突变造成的异常剪接模式
2026年1月 研究[J]. 南京医科大学学报(自然科学版),2026,46(1):55-67 · 61 ·
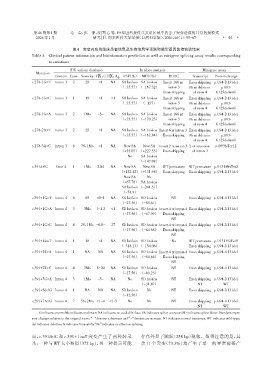
表4 突变对应的临床患者信息及生物信息学预测和微型基因剪接实验结果
Table 4 Clinical patient information and bioinformatics prediction as well as minigene splicing assay results corresponding
to variations
FⅨ variant database In silico analysis Minigene assay
Mutation
Domain Case Severity FⅨ:C FⅨ:Ag HSF(%) MES(%) RDDC Transcript Protein change
c.278⁃1G>T Intron 3 2 2S <1 NA SA broken SA broken Insert 168 nt Exon skipping p.G94⁃D131del
(-35.57)(-167.32) intron 3 96 nt deletion p.D93⁃
Exon skipping of exon 4 G125delinsG
c.278⁃1G>C Intron 3 1 1S <1 <1 SA broken SA broken Insert 168 nt Exon skipping p.G94⁃D131del
(-35.57) (-157) intron 3 96 nt deletion p.D93⁃
Exon skipping of exon 4 G125delinsG
c.278⁃1G>A Intron 3 2 1Mo -3- NA SA broken SA broken Insert 168 nt Exon skipping p.G94⁃D131del
(-35.57)(-170.23) intron 3 96 nt deletion p.D93⁃
Exon skipping of exon 4 G125delinsG
c.278⁃2A>T Intron 3 2 2S <1 NA SA broken SA broken Insert 6 nt intron 3 Exon skipping p.G94⁃D131del
(-35.57)(-162.84) Exonskipping 96 nt deletion p.D93⁃
of exon 4 G125delinsG
c.278⁃3A>G Intron 3 8 7S,1Mo <1 NA New SA New SA Insert 2 nt intron 3 2 nt retention p.D93fsTer12
(+55.07)(+227.53) Exonskipping of intron 3
No SA broken
(-147.08)
c.391delG Exon 4 1 1Mo 2.04 NA New SA New SA WT premature WT premature p.D131MfsTer2
(+122.42)(+131.98) Exon skipping Exon skipping p.G94⁃D131del
New SA No
(+67.78) SA broken
SA broken (-241.51)
(-74.9)
c.391+1G>T Intron 4 6 6S <0-1 NA SD broken SD broken NT Exon skipping p.G94⁃D131del
(-27.56) (-85.61)
c.391+1G>A Intron 4 3 3Mo 1-1.2 <1 SD broken SD broken Insert 4 nt intron 4 Exon skipping p.G94⁃D131del
(-27.56) (-87.99) Exonskipping
NT
c.391+1G>C Intron 4 4 2S,1Mo -0.8- 27 SD broken SD broken Insert 4 nt intron 4 Exon skipping p.G94⁃D131del
(-27.56) (-84.68) Exonskipping
NT
c.391+1insT Intron 4 1 1S <1 NA SD broken SD broken No WT premature p.D131VfsTer5
(-318.13)(-154.06) Exon skipping p.G94⁃D131del
c.391+2T>A Intron 4 1 NA NA NA SD broken SD broken Insert 4 nt intron 4 Exon skipping p.G94⁃D131del
(-27.56) (-84.68) Exonskipping
NT
c.391+2T>C Intron 4 4 3Mo 8-20 NA SD broken SD broken NT Exon skipping p.G94⁃D131del
(-27.56) (-80.23)
c.391+5G>A Intron 4 3 1Mo -5- NA No SD broken NT Exon skipping p.G94⁃D131del
(-31.47) NT WT
c.391+5delG Intron 4 1 NA NA NA SD broken No NT Exon skipping p.G94⁃D131del
(-12.56)
c.391+7A>G Intron 4 7 5S,2Mo <1-4 <1⁃3 No No NT Exon skipping p.G94⁃D131del
NT WT
S indicates severe;Mo indicates moderate;NA indicates no available data;SA indicates splice acceptor;SD indicates splice donor. Numbers repre⁃
sent changes relative to the original score;“-”denotes a decrease and“+”denotes an increase. NT indicates normal transcript;WT indicates wild⁃type;
del indicates deletion;fs indicates frameshift;“No”indicates no effect on splicing.
而,c.391delG 和 c.391+1insT 突变产生了两种转录 存在外显子跳跃(258 bp)现象。值得注意的是,其
本:一种与 WT 大小相似(372 bp),另一种提示可能 余 11 个突变(73.3%)均产生了单一的异常剪接产